Takashi Nakazawa, Professor at Nara Women’s University explores some fascinating aspects of chemistry and the archaeology of collagen, as well as a view point expressed on analysing ancient specimens in a collaborative way
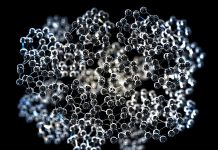
Collagen is the most abundant protein occurring in the body of mammals. There are nearly 30 types of collagen, the majority of which are fibrous. Type I collagen has a characteristic architecture of a triple helix consisting of polypeptide chains of two α1 and one α2. This naturally occurring biopolymer is ubiquitous in bones, teeth, and skins of animals, taking advantage of its fibrous structure. The triple helix of collagen is so resistant to ageing that it can often be found in archaeological specimens as old as tens of thousands of years. Consequently, it informs us of the animal species in terms of the respective amino acid sequence, even in the absence of DNA encoding genetic information. We are using mass spectrometry to obtain information of archaeological interest.
Chemistry
As well as its triple-helical structure, collagen has a few unique features including the primary structure such that glycine (G) appears every third residue in the sequence (G-X1-X2)n (n > 300). Moreover, the positions X1 and X2 are quite frequently occupied by proline (P) or hydroxyproline (many authors abbreviate this residue as O). The post-translational modification of P to O is almost specific to collagen and catalysed by proline hydroxylase just before the formation of the triple helix. It is widely accepted that the content and position of O is closely related to the stability of the collagen triple helix1. Before beginning the present study of collagen in archaeological specimens, we were investigating the correlation between the role of O and the stability of the triple helix in solution using collagen model peptides2. Since this modification occurs enzymatically, the position of O in the amino acid sequence is not defined in the genetic code, it is very difficult to determine which P residues are modified unless protein sequencing such as Edman degradation or mass spectrometry is conducted on collagen samples.
Archaeology
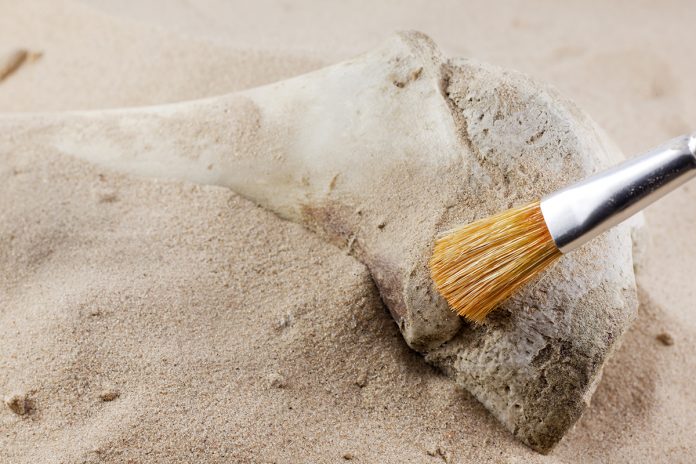
Our project of protein archaeology with mass spectrometry began about 10 years ago, when we detected cow collagen in Chinese ink stick excavated from the oldest Japanese capital palace site in Nara (Heijyo-Kyo) of the mid-sixth Century AD3. Encouraged by such an unexpected success of finding collagen as waste disposal kept in the soil as long as 1,250 years ago, we extended the list of proteins by seeking wider area from Nara to the sites worldwide and older ages. Actually, the age of our specimens became older from the binding media of Egyptian Romano portraits (180-200 AD)4, Egyptian wall paintings (2,400 BC) to West Asian Neolithic animal bones (5,000-8000 BC). And quite recently, we have been trying to detect collagen in Palaeolithic animal bones (30,000-35,000 BC). It is not surprising that the difficulty of analysis increases as the age of the specimens become older5. Nevertheless, we are excited about the challenge of solving this problem by mass spectrometry with the aid of protein chemistry6.
Note that the average lifetime of collagen to survive in archaeological specimens had been estimated to be much less than one million years7. It was, therefore, surprising that peptide fragments derived from collagen were found in the fossils of 8-million-year-old dinosaur bones8. However, another group has shown the complete match of amino acid sequences of collagen between that of modern ostrich (Struthio camelus) bone and those reported as of Tyrannosaurus and Brachylophosaurus, suggesting that there remains the possibility of cross-contamination of collagen9. In any case, we need to allow for the longevity of collagen, especially in the study of Palaeolithic bones for the identification of animal species.
Collaboration
Basically, we are analysing those ancient specimens in the collaborative study with archaeologists, scientists working for the conservation of cultural properties, artists, and historians, all of those who are least likely to work within our Laboratory of Biochemistry and Organic Chemistry in Nara Women’s University. One of these collaborations includes a project “Culture History of Paleo Asia” organised by Professor Yoshihiro Nishiaki (the University of Tokyo), supported by a Grant-in-Aid for Scientific Research on Innovative Areas (Grant No. 1802) from the Japanese Ministry of Education, Science, Culture, and Technology. In this project, we could distinguish between a goat and sheep as the species of Neolithic bones5. Without these collaborations and financial aids (see “Acknowledgement”), we could not do anything so exciting as to obtain a variety of archaeological materials needed to “read” a history written in terms of the chemical structure of collagen. For this project, we welcome researchers all over the world to collaborate with.
Acknowledgement
The author thanks the grants, 16H05656 and 17H05130, from the Minister of Education, Science, Culture, and Technology of Japan for financial aid.
References
1 Baum, J., Brodsky, B. (2000) Chapter 12, Case study 2: Folding of the collagen triple helix and its naturally occurring mutants. In Mechanism of Protein Folding. 2nd ed., Pain, R. H. ed., Oxford Univ. Press, Oxford.
2 Doi, M. et al. (2003) Characterization of collagen model peptides containing 4-fluoroproline; (4(S)-fluoroproline-Pro-Gly)10 forms triple helix but (4(R)-fluoroproline-Pro-Gly)10 does not. J. Am. Chem. Soc. 125, 9922-9923.
3 Kawahara, K. et al. (2011) Identification of animal species by the MALDI-MS of collagen in animal glues of Chinese ink sticks. Proceedings of 59th ASMS Conference and Allied Topics, Denver, CO, USA.
4 Mazurek, J. et al. (2014) Characterization of binding media in Egyptian Romano portraits using enzyme-linked immunosorbent assay and mass spectrometry. e-Preservation Sci. 11, 76-83.
5 Nakazawa, T. et al. (2018) Mass Spectrometry of Collagen Preserved in Neolithic Animal Bones for the Identification of Species. Proceedings of 66th ASMS Conference and Allied Topics, San Diego, CA, USA.
6 Nakazawa, T. et al. (2008) Terminal proteomics: N- and C-terminal analyses for high-fidelity identification of proteins using mass spectrometry. Proteomics 8, 673-685.
7 C. Nielsen-Marsh, (2002) Biomolecules in fossil remains. The Biochemist 24; 12-14.
8 Schweitzer, M. H. et al. (2009) Biomolecular characterization and protein sequences of the Campanian hadrosaur B. canadensis. Science 324: 626-631.
9 Buckley, M. et al. (2017) A fossil protein chimera; difficulties in discriminating dinosaur peptide sequences from modern cross-contamination. Proc. R. Soc. B 284: 20170544.
Please note: This is a commercial profile
Takashi Nakazawa
Professor
Nara Women’s University
Tel: +81 742 20 3396