Associate Professor of Medical Genetics, Silvia Deaglio, highlights the research strides being made in the fight against chronic lymphocytic leukemia
Next generation sequencing and functional genomic studies have contributed to bring about a radical change in our approach to the diagnosis and treatment of cancer. With the identification of recurrently mutated genes and constitutively activated signaling pathways, therapeutic approaches in oncology are becoming highly specific and personalised with the aim of maximising results and minimising side effects.
Chronic lymphocytic leukemia
Chronic lymphocytic leukemia (CLL) is an outstanding example in this context. This disease, which represents the most common form of adult leukemia in Europe and North America, has now several highly effective targeted therapeutic options that are rapidly substituting the “old” regimes of chemo and immunotherapy. The most advanced clinical experience has been accumulated using the btk inhibitor ibrutinib, which targets a specific kinase in the signaling pathway activated by the B cell receptor, clearly the driving force in the progression of the disease. Even if clinical trials using btk inhibitors are proving highly successful, with very few cases progressing under the drug and with durable responses even in the case of patients with unfavorable genetic aberrations, the cure is still out of the picture. Importantly, some studies have suggested that discontinuation of the drug leads to rapid disease relapse and progression, posing questions on the long-term effects of these therapies and also on its costs.
The Immunogenetics Unit of the Human Genetics Foundation, an institution that opened in Turin at the end of 2010 from the joint efforts of the local University, the Polytechnic and the Compagnia di SanPaolo, has been working since the beginning in the CLL field, with the aim of unraveling pathways and mechanisms controlling leukemia proliferation and survival. These studies, significantly supported by the Italian Association for Cancer Research (AIRC), have contributed to our knowledge on how the CLL cell grows and how it manipulates its microenvironment to create local conditions that favor growth and survival.
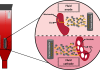
One of the areas of research that has attracted our attention as potentially interesting from the therapeutic point of view is based on data indicating that leukemic cells need to be surrounded and supported by non-tumor cells to survive and proliferate. These dangerous liaisons take place in privileged niches within the lymph nodes known as proliferation centers (Figure 1). Our group has shown that extracellular adenosine is one of the means through which CLL cells obtain T and myeloid cell cooperation. This small molecule is the final product of extracellular ATP metabolism, generated through enzymes expressed on the surface of both CLL and non-leukemic cells, and signals through specific membrane receptors. We showed that adenosine effects on CLL homeostasis are twofold: on one side, it directly protects leukemic cells from spontaneous and drug-induced apoptosis; on the other side, it shapes the environment towards immune tolerance, by inhibiting T-cell responses and inducing differentiation of supportive myeloid cells (Serra et al., 2011; Serra et al., 2016).
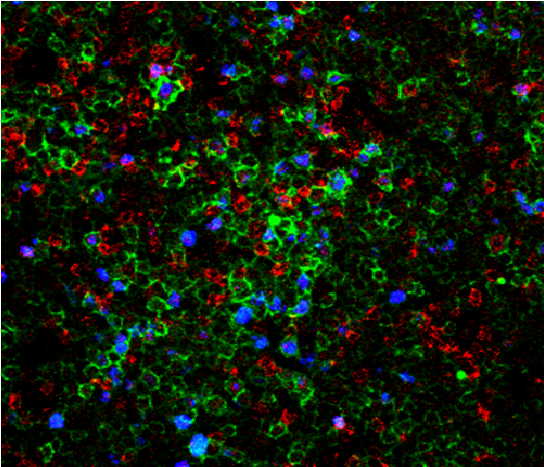
In addition to tricks adopted to mold a tumor-friendly environment, CLL cells exhibit intrinsic features that heavily impact their behavior, and whose functional characterisation represents the second main topic of our research activity. Before the advent of next generation sequencing, there were few markers to identify patients with an aggressive disease and the expression of CD38 was one of them. Our group has extensively investigated the role and the signaling properties of CD38 and revealed that it actively favors survival and proliferation of leukemic cells by cooperating with chemokine receptors and integrin’s. By blocking CD38 enzymatic activity, we also provided proof of the principle that drugs or monoclonal antibodies targeting this molecule might be therapeutically useful in CLL (Vaisitti et al., 2015).
More recently, the use of ultra-deep sequencing platforms, led to the identification of recurrent mutations associated with worse disease outcomes and likely conferring advantages to leukemic cells. These platforms are also used to monitor disease evolution, clearly showing that the genetic profile of CLL evolves with time and therapy. This observation implies that either leukemic cells acquire novel genetic lesions as a result of micro -environmental conditions that lead to genomic instability, or that sub clones of the leukemic population, carrying specific mutations are progressively selected during the course of the disease. This discovery pointed out the importance of understanding and predicting the effects exerted by mutated genes in the disease context and how they affect leukemic cell behavior and therapy responses. Our team has focused on NOTCH1 mutations, as they are relatively frequent in newly diagnosed CLL, increasing with the progression of the disease and in conditions of therapy resistance. We have shown that NOTCH1 mutations deregulate its activities as a transcription factor, remodeling gene expression of leukemic cells through direct (transcription-mediated) or indirect (epigenetic) mechanisms. Functionally, mutated NOTCH1 promotes growth and migration of leukemic cells by increasing expression of the chemokine receptor CCR7, through a complex network of nuclear signaling. As a result, mutated cells exhibit increased chemotaxis toward CCL19, the ligand of CCR7 and an important chemokine regulating trafficking of CLL cells to lymphoid niches. The relevance for the disease is that NOTCH1 mutations may contribute to an unfavorable prognosis by promoting the migration of leukemic cells to the lymphoid organs, where the protective micro-environment provides stimulatory signals favoring a more aggressive behaviour (Arruga et al., 2016).
Even if it is difficult to predict the future of this highly interesting and rapidly evolving field, the shared hope is to obtain a cure suitable for the majority of patients. It is possible that this will come by combining drugs that target central pathways in leukemic cell biology together with drugs that alter connections and communications with the environment, making it inhospitable. The next decade will likely tell if these approaches work.
References
Arruga, F., B. Gizdic, C. Bologna, S. Cignetto, R. Buonincontri, S. Serra, T. Vaisitti, K. Gizzi, N. Vitale, G. Garaffo, E. Mereu, F. Diop, F. Neri, D. Incarnato, M. Coscia, J. Allan, R. Piva, S. Oliviero, R.R. Furman, D. Rossi, G. Gaidano, and S. Deaglio. 2016. Mutations in NOTCH1 PEST-domain orchestrate CCL19-driven homing of Chronic Lymphocytic Leukemia cells by modulating the tumor suppressor gene DUSP22. Leukemia.
Serra, S., A.L. Horenstein, T. Vaisitti, D. Brusa, D. Rossi, L. Laurenti, G. D’Arena, M. Coscia, C. Tripodo, G. Inghirami, S.C. Robson, G. Gaidano, F. Malavasi, and S. Deaglio. 2011. CD73-generated extracellular adenosine in chronic lymphocytic leukemia creates local conditions counteracting drug-induced cell death. Blood 118:6141-6152.
Serra, S., T. Vaisitti, V. Audrito, C. Bologna, R. Buonincontri, S.S. Chen, F. Arruga, D. Brusa, M. Coscia, O. Jaksic, G. Inghirami, D. Rossi, R.R. Furman, S.C. Robson, G. Gaidano, N. Chiorazzi, and S. Deaglio. 2016. Adenosine signaling mediates hypoxic responses in the chronic lymphocytic leukemiamicroenvironment. Blood Advances 1:47-61.
Vaisitti, T., V. Audrito, S. Serra, R. Buonincontri, G. Sociali, E. Mannino, A. Pagnani, A. Zucchetto, E. Tissino, C. Vitale, M. Coscia, C. Usai, C. Pepper, V. Gattei, S. Bruzzone, and S. Deaglio. 2015. The enzymatic activities of CD38 enhance CLL growth and trafficking: implications for therapeutic targeting. Leukemia 29:356-368.
Silvia Deaglio, MD, PhD
Associate Professor of Medical Genetics
University of Turin & Immunogenetics Unit, Human Genetics Foundation
silvia.deaglio@unito.it
Please note: this is a commercial profile